Width Helps and Hinders Splitting Flows
Jan 1, 2024·
 ,
,
,
,
,
,
,
·
0 min read
,
,
,
,
,
,
,
·
0 min read
Manuel Cáceres
Massimo Cairo
Andreas Grigorjew
Shahbaz Khan
Brendan Mumey
Romeo Rizzi
Alexandru I. Tomescu
Lucia Williams
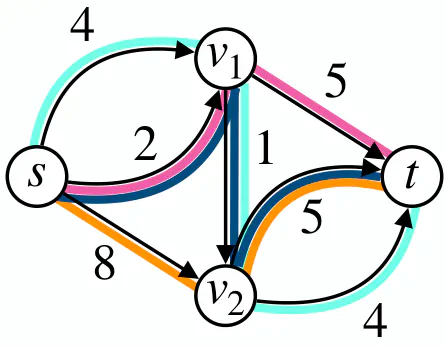 Flow decomposition using negative weights can yield to a smaller solution
Flow decomposition using negative weights can yield to a smaller solution
Abstract
Minimum flow decomposition (MFD) is the NP-hard problem of finding a smallest decomposition of a network flow/circulation $X$ on a directed graph $G$ into weighted source-to-sink paths whose weighted sum equals $X$. We show that, for acyclic graphs, considering the width of the graph (the minimum number of paths needed to cover all of its edges) yields advances in our understanding of its approximability. For the version of the problem that uses only non-negative weights, we identify and characterise a new class of width-stable graphs, for which a popular heuristic is a $O(\log Val(X))$-approximation ($Val(X)$ being the total flow of $X$), and strengthen its worst-case approximation ratio from $\Omega(\sqrt{m})$ to $\Omega(m/\log{m})$ for sparse graphs, where $m$ is the number of edges in the graph. We also study a new problem on graphs with cycles, Minimum Cost Circulation Decomposition (MCCD), and show that it generalises MFD through a simple reduction. For the version allowing also negative weights, we give a $(\lceil \log ||X|| \rceil + 1)$-approximation ($||X||$ being the maximum absolute value of $X$ on any edge) using a power-of-two approach, combined with parity fixing arguments and a decomposition of unitary circulations ($||X||\le 1$), using a generalised notion of width for this problem. Finally, we disprove a conjecture about the linear independence of minimum (non-negative) flow decompositions posed by [Kloster et al. 2018], but show that its useful implication (polynomial-time assignments of weights to a given set of paths to decompose a flow) holds for the negative version.
Type
Publication
In ACM Transactions on Algorithms