Fast Indexes for Gapped Pattern Matching
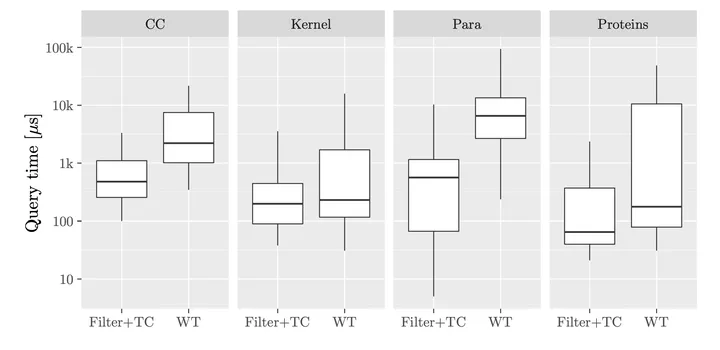 Performance of our method versus wavelet trees
Performance of our method versus wavelet trees
Abstract
We describe indexes for searching large data sets for variable-length-gapped (VLG) patterns. VLG patterns are composed of two or more subpatterns, between each adjacent pair of which is a gap-constraint specifying upper and lower bounds on the distance allowed between subpatterns. VLG patterns have numerous applications in computational biology (motif search), information retrieval (e.g., for language models, snippet generation, machine translation) and capture a useful subclass of the regular expressions commonly used in practice for searching source code. Our best approach provides search speeds several times faster than prior art across a broad range of patterns and texts.
Type
Publication
In SOFSEM 2020