Faster Repetition-Aware Compressed Suffix Trees Based on Block Trees
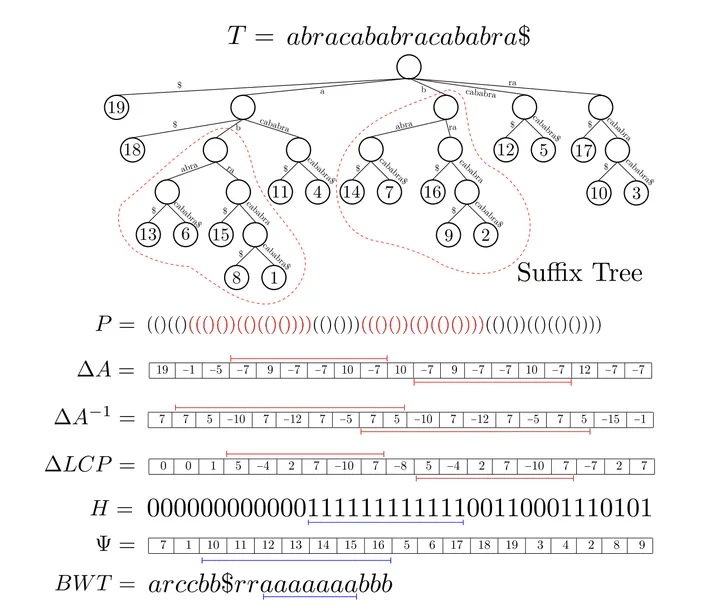 CST concepts
CST concepts
Abstract
Suffix trees are a fundamental data structure in stringology, but their space usage, though linear, is an important problem in applications. We design and implement a new compressed suffix tree targeted to highly repetitive texts, such as large genomic collections of the same species. Our suffix tree builds on Block Trees, a recent Lempel-Ziv-bounded data structure that captures the repetitiveness of its input. We use Block Trees to compress the topology of the suffix tree, and augment the Block Tree nodes with data that speeds up suffix tree navigation.
Our compressed suffix tree is slightly larger than previous repetition-aware suffix trees based on grammars, but outperforms them in time, often by orders of magnitude. The component that represents the tree topology achieves a speed comparable to that of general-purpose compressed trees, while using 2–10 times less space, and might be of independent interest.
Type
Publication
In SPIRE 2019